Challenges & Rewards
Every few years we poll our community of biology educators to find out what topics they’re finding particularly tricky to teach and to help us figure out what content to develop. We then consider where our “special sauce” (simulated experiments and other engaging interactives) will be most effectively applied. DNA replication, transcription & translation, and gene regulation regularly make up the “top ten trickiest topics to teach” list. This complexity presented unique challenges for developing discovery-based learning activities for our new Molecular Biology Series. Good thing we enjoy challenges!
Background
The fields of ecology and evolutionary biology have strong traditions of mathematics and simulations that inform research. The teaching tools upon which SimBio was founded over 25 years ago drew on these research traditions to develop suites of virtual labs, and later ecology and evolutionary biology tutorials, that let students explore and “discover” fundamental biological concepts.
My own research background is more molecular, and I always dreamed of taking the philosophy of learning via exploration and discovery we pioneered with our ecology and evolution labs and applying it to cellular/molecular level processes. However, with a few exceptions (such as Action Potentials), cellular and molecular biology do not share the deep research base of mathematics and simulations found in ecology and evolution. Given the current state of molecular biology knowledge, we often teach undergraduates “just-so” stories about ideas – such as how information flows from DNA to protein or how cells store and use energy. Learning these fields includes understanding complicated “minutia” (e.g., functions of specific proteins and interactions between various molecules), which can be overwhelming and impede diving deeper into underlying mechanisms. Our challenge is to figure out how to make that important learning an engaging, inquiry-driven experience.
Innovations in active learning
Led by our molecular biology author Kerry Kim, and after learning from failed attempts and iterations in our design, our team developed engaging approaches to help students understand key concepts in molecular biology. As an example, here is a simulation of DNA replication from DNA Explored.
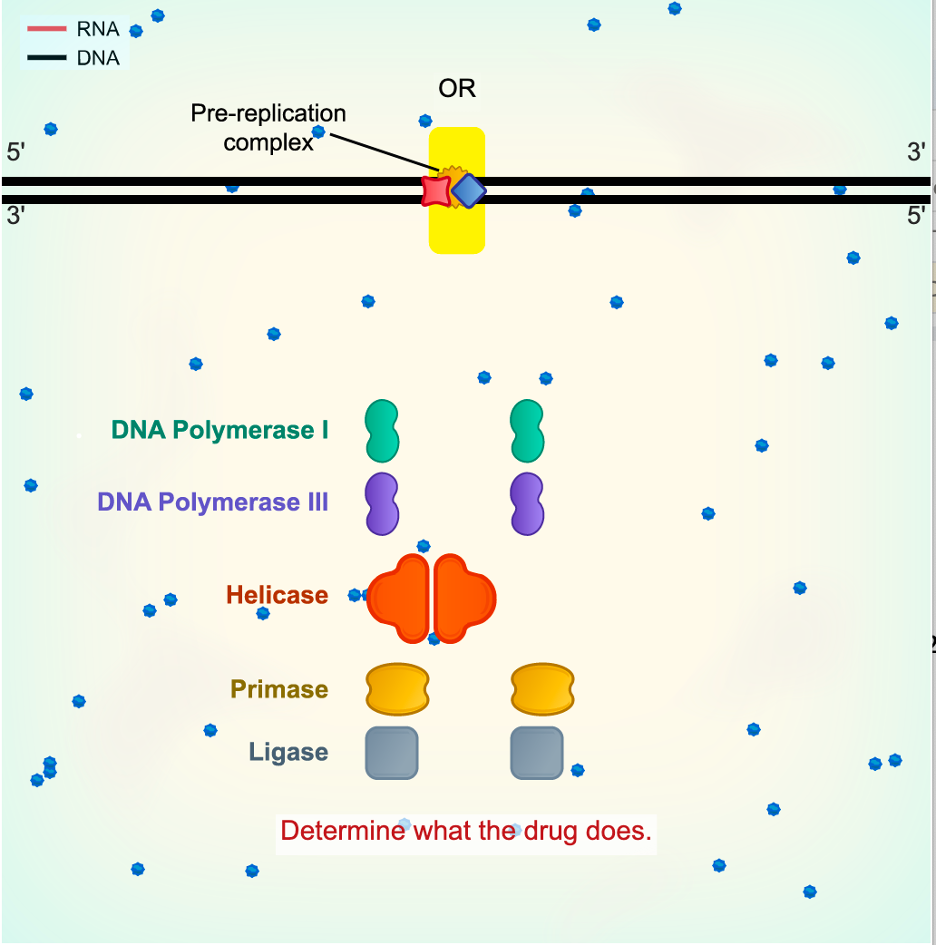
It includes many of the enzymes involved in replication. Students use these to replicate the DNA at the top by dragging them in the appropriate order and to the right place. The underlying simulation shows what happens if you get something wrong. For instance, if DNA Polymerase is placed on the DNA without an RNA primer, it will fall off. Primase won’t make RNA primer if the DNA strands are not first separated, and so on. Pop-up feedback helps students who get stuck. In this way, a student not only memorizes the order of events, but through their own experimentation sees why things happen in that order. They then get to apply that experimental thinking, in the simulation shown above, to figure out how an anti-cancer drug works.
Using similar approaches, our latest molecular biology modules explore how information flows in cells, starting with DNA structure and replication, then Transcription and Translation, and, just released this winter, a tutorial/lab on Gene Regulation. These add to our growing collection of cell biology tutorial/labs covering respiration, diffusion, osmosis, action potentials, Mendelian Pigs genetics, experimental design and more. A growing number of Intro Bio classes are replacing their traditional textbooks with our active-learning-based tutorial/labs, which demonstrably help students learn complex topics. I’m hopeful that the opportunities to tinker with molecular systems is also helping students understand and appreciate their logic and elegance.
My colleague Kerry Kim, periodically gives webinars demonstrating the new Molecular Biology Series. I always love showing them as well, and you can try them yourself through our evaluation software. If you’d like to see how we help students explore the molecular basis of biology, please get in touch.






 Helping Students Become Better Graphers
Helping Students Become Better Graphers

